DL-QC-fNIRS: A Deep Learning Tool for Automated fNIRS Signal Quality Control
A new deep learning framework from the University of Zurich's Biomedical Optics Research Lab uses convolutional neural networks to automatically assess fNIRS signal quality channel by channel, achieving over 93% accuracy and outperforming traditional metrics like CV and SCI.
A new study published in Neurophotonics by Guglielmini, Chen, and Wolf (University Hospital Zurich, University of Zurich, and ETH Zurich) introduces DL-QC-fNIRS — a deep learning framework that automates one of the most tedious and error-prone steps in fNIRS preprocessing: channel-wise signal quality assessment.
Why Signal Quality Control Matters in fNIRS
In functional near-infrared spectroscopy (fNIRS) research, ensuring signal quality is a critical preprocessing step. Bad channels — those contaminated by poor optode coupling, motion, or extracerebral noise — can derail downstream statistical analyses if not identified and excluded. Traditionally, researchers have relied on index-based metrics such as the coefficient of variation (CV) and the scalp coupling index (SCI) to flag noisy channels.
The problem is that these metrics depend on arbitrary thresholds. A channel that scrapes by the SCI cutoff in one study may be rejected in another, and both the CV and SCI frequently misclassify channels — keeping noisy ones and discarding usable ones. This introduces variability across labs and reduces the reproducibility of fNIRS findings.
The DL-QC-fNIRS Approach
The authors take a different angle. Instead of relying on summary statistics, they convert each channel's oxyhemoglobin time series into a continuous wavelet transform (CWT) scalogram — a 2D time-frequency representation of the signal — and feed that scalogram into a convolutional neural network (CNN) trained to classify it as good or bad.
To improve physiological specificity, the pipeline also extracts subject-specific cardiac frequency information. Because clean fNIRS signals carry a clear cardiac pulsation component, the model can use the presence or absence of that signature as a strong indicator of channel quality.
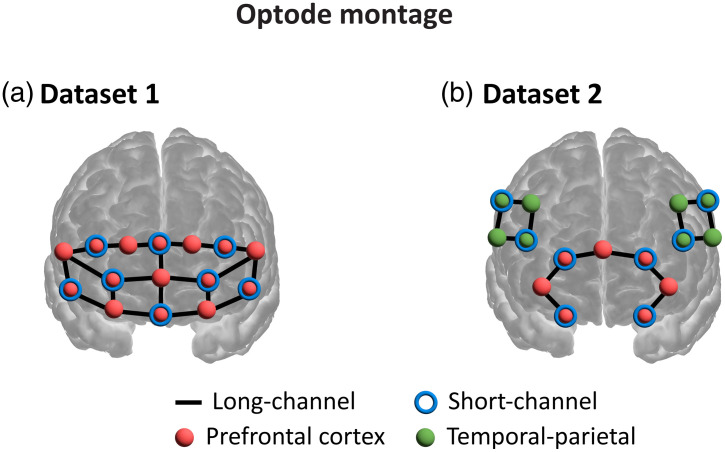
The team benchmarked four CNN architectures — GoogLeNet, ResNet-50, SqueezeNet, and EfficientNet-B0 — on two independent datasets and one combined heterogeneous dataset, to see which architecture generalizes best across recording conditions.
Results: GoogLeNet Wins, Deep Learning Beats Index Metrics
GoogLeNet achieved the highest accuracy on the combined dataset, exceeding 93%, with strong sensitivity and specificity across test sets. More importantly, when compared head-to-head with the CV and SCI metrics, DL-QC-fNIRS produced markedly higher F1-scores and a more favorable balance between sensitivity and specificity — meaning fewer noisy channels slip through and fewer good channels get unnecessarily rejected.
The performance held up across the heterogeneous dataset, suggesting the model is robust to differences in hardware, montage, and subject population.
Open-Source MATLAB GUI
One of the most practically useful aspects of this work: the authors released DL-QC-fNIRS as an open-source, MATLAB-based graphical interface, available on GitHub. The tool supports two workflows — applying a pretrained model to your dataset, or training a new model on your own labelled data through the GUI. The code is publicly available at github.com/zhuofeichen312/DL-QC-fNIRS.
That accessibility matters. One of the persistent barriers to adopting machine learning methods in fNIRS labs has been the need for custom Python pipelines and ML expertise. By packaging the framework as a GUI with built-in pretrained models, the authors lower the barrier to entry and make standardized, reproducible quality control practical for any fNIRS group.
What This Means for fNIRS Research
Quality control has long been a soft underbelly of the fNIRS pipeline — every lab has its own thresholds, every reviewer has their own concerns, and disagreements about which channels to keep can swing results. A standardized, validated, automated tool changes that conversation. It moves quality control from a judgment call into a reproducible step that other labs can replicate exactly.
For neuroscience and clinical fNIRS researchers using systems like NIRx, Artinis, or any compatible platform, DL-QC-fNIRS offers a way to standardize the preprocessing pipeline, reduce reviewer-induced variability, and ultimately strengthen the reproducibility of fNIRS findings.
Reference
Guglielmini, S., Chen, Z., & Wolf, M. (2025). DL-QC-fNIRS: a deep learning tool for automated quality control in functional near-infrared spectroscopy signals. Neurophotonics, 13(1), 015001. https://doi.org/10.1117/1.NPh.13.1.015001

NEED MORE INFORMATION ABOUT THIS PRODUCT?
Send us your emailAdvance Your Research
Contact NBT today for expert consultation on your neuroscience instrumentation needs.
.svg)
.jpg)


.svg)

